Analysis for Disease Gene Association Using Machine Learning
Objective
In this project, we propose and analyze a novel computational methods for the identification of genes associated with diseases using Machine Learning techniques.
Abstract
To recognize the basis of disease, it is essential to determine its underlying genes. Understanding the association between underlying genes and genetic disease. This is a fundamental problem regarding human health. Identification and association of genes with the disease require time consuming and expensive experimentations of a great number of potential candidate genes.
Therefore, the alternative inexpensive and rapid computational methods have been proposed that can identify the candidate gene associated with a disease. Most of these methods use phenotypic similarities due to the fact that genes causing same or similar diseases have less variation in their sequence or network properties of proteinprotein interactions based on-premises that genes lie closer in protein interaction network that causes the similar or same disease.
In this project, we propose and analyze a novel computational methods for the identification of genes associated with diseases. Some advance topological and biological features that are overlooked currently are introducing for identifying candidate genes. We evaluate different computational methods on disease-gene association data from DisGeNET in a 10-fold cross-validation mode based on TP rate, FP rate, precision, recall, F-measure, and ROC curve evaluation parameters.
Keywords: Disease Gene Association, Protein-Protein Interaction Network (PPIN), Electron-Ion Interaction Pseudopotential (EIIP).
NOTE: Without the concern of our team, please don't submit to the college. This Abstract varies based on student requirements.
Block Diagram
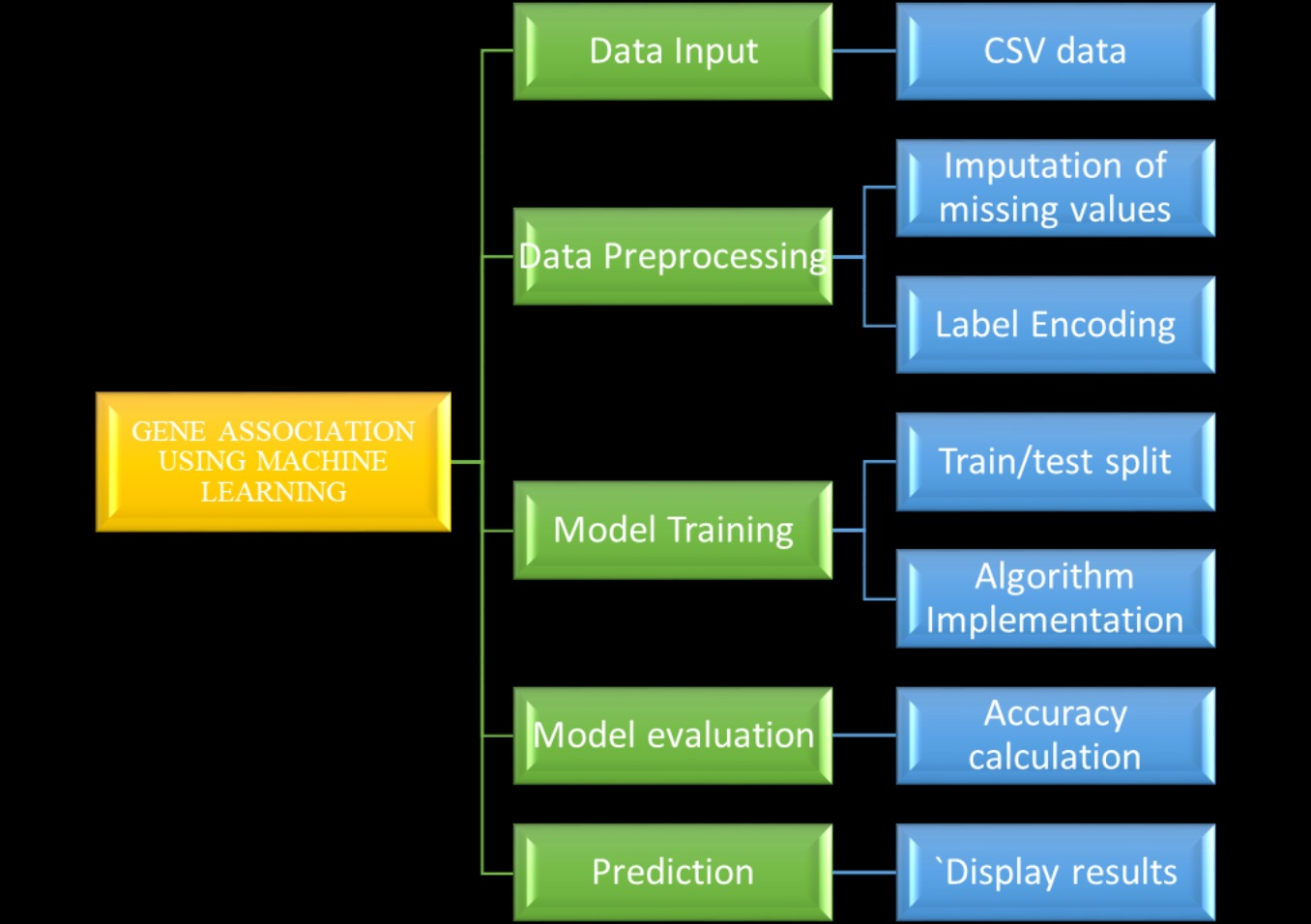
Specifications
HARDWARE SPECIFICATIONS:
- Processor- I3/Intel Processor
- RAM- 4GB (min)
- Hard Disk- 128 GB
- Key Board-Standard Window
- Keyboard. Mouse-Two or Three Button Mouse.
- Monitor-Any.
SOFTWARE SPECIFICATIONS:
- Operating System: Windows 7+
- Technology: Python 3.6+
- IDE: PyCharm IDE
- Libraries Used: Pandas, NumPy, Scikit-Learn, Matplotlib.
Learning Outcomes
- Scope of Real Time Application Scenarios.
- Objective of the project .
- How Internet Works.
- What is a search engine and how browser can work.
- What type of technology versions are used.
- Use of HTML , and CSS on UI Designs.
- Data Parsing Front-End to Back-End.
- Working Procedure.
- Introduction to basic technologies used for.
- How project works.
- Input and Output modules.
- Frame work use.
- Datasets properties.
- Machine learning algorithms.
- Data preprocessing techniques.
- 10 fold cross techniques.
- Underlying and genetic diseases.
- What are biological and topological features set.
- Graphs for drawing based on highest accuracy model.
- Project Development Skills:
- Problem analyzing skills.
- Problem solving skills.
- Creativity and imaginary skills.
- Programming skills.
- Deployment.
- Testing skills.
- Debugging skills.
- Project presentation skills.
- Thesis writing skills.





 Paper Publishing
Paper Publishing
